-
Notifications
You must be signed in to change notification settings - Fork 2
Example extractions
Please see the example extractions below for some further clarification on the extraction process. Note that we extract data as written in the article, even if it appears that there are errors (terminology or otherwise). In this case, extract as written, leave a note on the article form and bear this in mind for the quality assessment.
This extraction is based on the example from training by Thom Rawson, using the following paper:
Xiao Z, Li Y, Chen R, Li S, Zhong S, Zhong N. A retrospective study of 78 patients with severe acute respiratory syndrome. Chin Med J (Engl). 2003 Jun;116(6):805-10. PMID: 12877784.
First, fill in your name, email address, and declare that you will perform extractions accurately (we will use Thom's name and email address for this example):

Then fill out key article detail:

This paper contains no models or outbreaks to extract, so tick the appropriate boxes:


The paper states: "A history of close contact was reported in 56 cases (71.8%), of whom, incubation periods were available in 17 cases (1-5 days, mean 2.4 $$\pm$$1.5 days)."
We can therefore extract the incubation period mean (2.4) and range (under variability, *1-5). We can also extract what we think is a measure of uncertainty (1.5), although the text does not describe exactly what measure this is:

The sample size for this parameter is also given (17), but we can search the paper for other contextual information as found in the introduction and methods:
"A patient from Guangdong province ..." - Introduction.
"A total of 78 cases with SARS in Guangdong were admitted to GIRD (Guangzhou Institute of Respiratory Diseases) between December 22, 2002 and near the end of March 2003 ... The patients comprised 42 men and 36 women, aged 20-75" - Methods.
This gives us contextual information, including sex (both male and female), total sample size (78), setting (Hospital based), group ("cases with SARS", implies "persons of interest"), age (2-75), location (Guangdong), location type (province), survey context (December 22, 2002 to an unknown date near the end of March 2003). Please note that the incubation period is based on a subset of 17 individuals and we do not have the age or sex breakdown for this subset, and cannot therefore extract these fields for this parameter. Nor do we have the day of the last sampling date, so we can extract only the month and year for this date (xx-03-2003).

...

The text says: "The hospital stay of the 78 cases was (23.1 +- 11.9) days (ranged 5-78 d)."
We can, again, extract the mean (23.1), range (5-78), and unknown uncertainty measure (11.9):

Note that we can now use the full sample size (78), with sex (both) and age (2-75) information:

"Seventy-one patients were cured and ... 9%". Some of the text cannot be seen here, but in the abstract it also says:
"Seven patients who developed ARDS complicated with multiple organs dysfunction syndrome (MODS) died."
From which we can (slightly) infer 7 died from a sample of 78 which is about 9%, and extract the CFR. Note that we can also extract the numerator and denominator, separately.

The sample context is the same as for parameter 2.
Table 1 identifies the significance of acute lung injury (ALI) according to underlying diseases using a chi-squared test. Table 2 similarly reports the significance of age and neutrophils as risk factors of ALI.
We can extract these as risk factors where we interpret ALI as the risk factor outcome (severe disease), and extract underlying disease (comorbidity), age, and neutrophils (other) as the risk factors. The tests performed were chi-squared tests and t-tests which are not adjusted tests.

We also want to extract the non-significant risk factors, but do so as a separate parameter.
Table 2 reports that lymphocytes are not a significant risk factor for ALI. We can extract this under other as follows.

Finally, we perform the quality assessment. We must consider the QA guidance for each parameter extracted (human delays, severity and risk factors) and decide if the QA questions can be answered for each of these extractions. The paper meets the criteria for all described, but does not examine possible biases in the data and therefore fails QA6 and QA7.

Your study states that R0 has mean 2.37 (95% CI = 1.92 – 2.54) and standard deviation 0.37 +/- 0.12 standard error. Our extraction tool allows us to extract the uncertainty around the mean (the 95% CI), the variability of the data (SD) and the uncertainty around the variability of the data (SE of the SD). You would extract these in a single parameter form, as follows:
- Parameter type: Reproduction number (Basic R0)
- Parameter value: 2.37
- Parameter value type: Mean
- Single: leave blank
-
Paired:
- Type: CI95%
- Lower value: 1.92
- Upper value: 2.54
- Would you like to add associated parameter?: Yes
- Parameter - value type: Standard Deviation
- Parameter – single value: 0.37
- Would you like to extract variability uncertainty?: Yes
-
Parameter uncertainty: Single
- Type: Standard Error (SE)
- Value: 0.12
Most instances of distributions to extract are straightforward and recorded using the distribution section, specifying the relevant distribution, "parameter A" and "parameter B". A more complicated instance is that of over-dispersion parameters as we need to capture slightly different amounts of information, depending on the underlying model.
E.g. 2a: Superspreading and the effect of individual variation on disease emergence [doi:10.1038/nature04153]: “For the outbreak, the maximum-likelihood estimate k^ is 0.16 (90% confidence interval 0.11–0.64), indicating an underlying distribution of n that is highly overdispersed”. The paper states that it uses a branching process model.
- Parameter type: Overdispersion
- Parameter value: 0.16
- Parameter value type: Mean
- Single: leave blank
-
Paired:
- Type: CI90%
- Lower value: 0.11
- Upper value: 0.64
- Method: Renewal equation/branching process.
E.g 2b: Modeling Heterogeneity in Direct Infectious Disease Transmission in a Compartmental Model [doi:10.3390/ijerph13030253]: This example is more complicated to extract. The paper states that it captures heterogeneity before and after 20th April, when non-pharmaceutical interventions (NPIs) were implemented, and which we capture in the parameter context section.
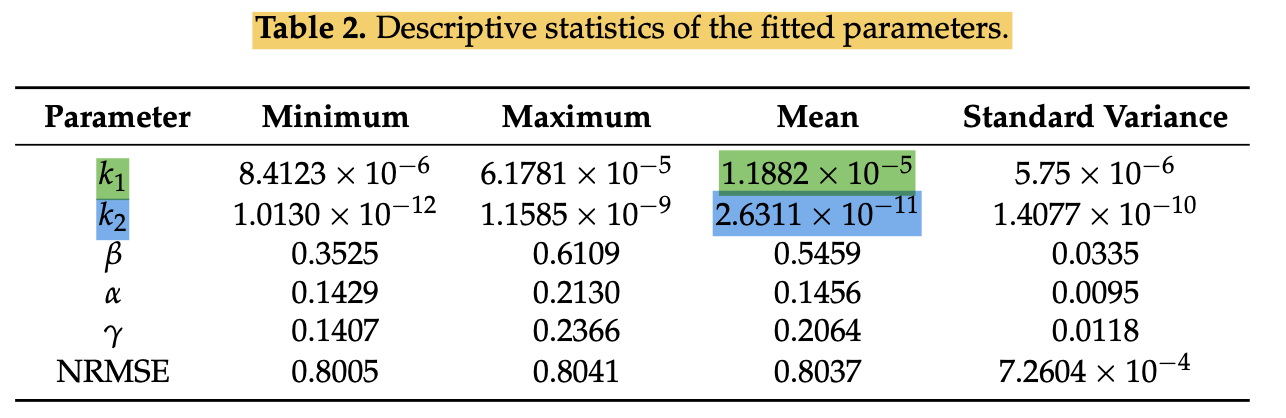
- Parameter type: Overdispersion
- Parameter value: 1.882
- Parameter exponent: -5
-
Parameter range of central estimates:
- Lower value: 8.4123
- Upper value: 6.1781
- Parameter value type: Mean
-
Single:
- Type: Standard Deviation
- Value: 5.75
-
Context – survey:
- Start date: 07-03-2003
- End date: 20-04-2003
Notes: Uncertainty Standard Deviation and Parameter Range lower value have an exponent of 10-6 (different from the mean which is 10-5).
- Parameter type: Overdispersion
- Parameter value: 2.6311
- Parameter exponent: -11
-
Parameter range:
- Lower value: 1.0130
- Upper value: 1.1585
- Parameter value type: Mean
-
Single:
- Type: Standard Deviation
- Value: 1.4077
- Method: Compartmental Model
-
Context – survey:
- Start date: 20-04-2003
- End date: 04-06-2003
Notes: Parameter Range and uncertainty values have different exponents from the mean (which is -11): Range Lower (-12), Upper (-9), Standard deviation (-10).
- Parameter type: Human delay – latent period
- Parameter value: 0.1456
-
Parameter range:
- Lower value: 0.1429
- Upper value: 0.2130
- Parameter value type: Mean
-
Single:
- Type: Standard Deviation
- Value: 0.0095
- Method: Compartmental Model.
- Parameter type: Human delay – infectious period
- Parameter value: 0.2064
-
Parameter range:
- Lower value: 0.1407
- Upper value: 0.2366
- Parameter value type: Mean
-
Single:
- Type: Standard Deviation
- Value: 0.0118
- Method: Compartmental Model.
E.g. 3a: Repeated serology: “Seroprevalence in 2003 was 4.2%. We repeated the serological survey in 2013, two years after the last outbreak, and found seroprevalence was 8.9%. All tests were conducted using IgG”. Here you would extract using two parameter forms to capture each seroprevalence estimate in turn.
- Parameter type: Seroprevalence - IgG
- Parameter value: 4.2
- Parameter value unit: Percentage (%)
Then we record the estimate-specific detail in the context section: e.g. sample size in 2003, and 2003 survey dates.
- Context – timing: pre-outbreak
- Parameter type: Seroprevalence - IgG
- Parameter value: 8.9
- Parameter value unit: Percentage (%)
Then record the estimate-specific detail in the context section: e.g. sample size in 2013, and 2013 survey dates.
- Context – timing: post-outbreak
E.g. 3b: Neutralisation tests: ELISA tests are prone to issues with cross-reactivity, and so depending on the pathogen, positive results may need to be confirmed via a neutralisation assay (e.g. plaque reduction neutralisation test (PRNT); pseudovirus neutralisation test (pPNT)). However, neutralisation tests are expensive and require specific equipment, and so often only all positive tests and a subset of the negative tests are sent for this testing. For example, consider a serosurvey in which there were 100 ELISA tests conducted of which 10 were positive. 20 samples - all 10 positive tests and 10 randomly-selected ELISA-negative tests - are tested via PRNT. This returns 5 positives overall, corresponding to a seroprevalence of 25%. You would extract this as follows:
- Parameter type: Seroprevalence – (whichever was the initial ELISA test)
- Parameter value: 10
- Parameter value unit: Percentage (%)
- Numerator: 10
- Denominator: 100
- Parameter type: Seroprevalence - PRNT
- Parameter value: 25
- Parameter value unit: Percentage (%)
- Numerator: 5
- Denominator: 20
Notes: Please use the “notes” box to provide further detail on this (e.g. copy and paste text from the paper regarding the neutralisation tests).
Based upon this paper which we used as an example for test extraction: https://journals.plos.org/plosntds/article?id=10.1371/journal.pntd.0004622 This was a survey of children of Ebola survivors. In this study they went house to house to look at how many children were infected and what their outcomes were. Here, in the parameter context the extraction would look as follows:
- Setting: Household based
- Group: Household contacts of survivors