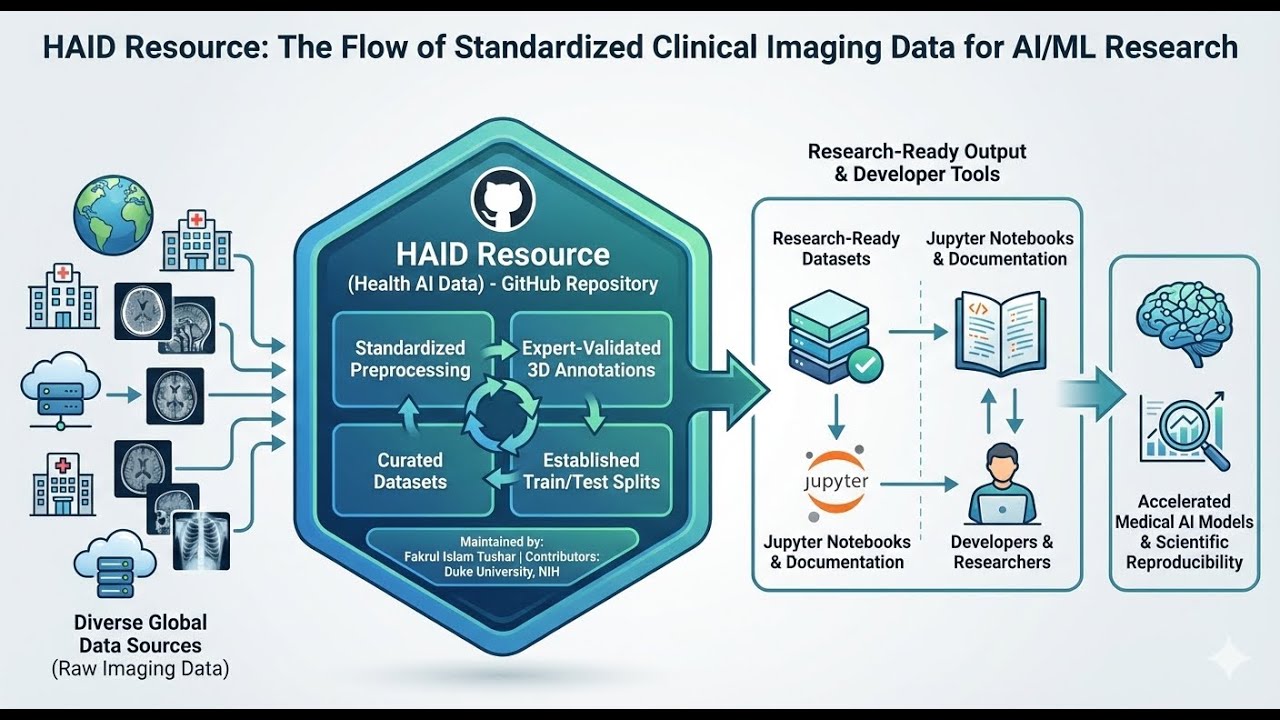
The HAID (Health AI Data Resource) is an open-access repository hosted on GitHub that provides standardized medical imaging datasets to support the development of artificial intelligence and machine learning models. Curated by Fakrul Islam Tushar, the platform currently offers 13 distinct clinical collections covering conditions such as lung cancer, COVID-19, and universal lesions. Each resource includes preprocessed imaging files, expert-verified annotations, and consistent train-test splits to ensure research reproducibility. Beyond raw data, the repository serves as a functional toolkit by supplying curated Jupyter notebooks and reference workflows for data analysis. By unifying diverse international datasets with comprehensive documentation, HAID aims to democratize high-quality clinical data for the global medical imaging community.
🚀 Interactive Research :You can chat with the source materials for this project using our NotebookLM Dashboard: 👉 Access the Interactive Notebook
📊 Slide Deck View the technical overview of the HAID project: 👉 Download Slide Deck (PDF)
📑 Resource Tables at a Glance
| 📂 Table | 📝 One-line Description |
|---|---|
| Available Datasets | Preprocessed, standardized medical imaging datasets with ready-to-use splits and documentation. |
| Associated Open-Access Tools & Studies | HAID-developed tools, benchmarks, and in-silico studies for detection, segmentation, and classification. |
| Community & Collaborator Open-Source Resources | External open-source tools and curated resources shared by collaborators and the community. |
🎥 HAID Intro Video
Click the image above to watch the introduction to HAID on YouTube
To democratize access to high-quality, preprocessed medical imaging datasets by providing:
- ✅ Standardized preprocessing pipelines
- ✅ Comprehensive documentation
- ✅ Ready-to-use annotations for detection, segmentation, and classification
- ✅ Train/validation/test splits for reproducible benchmarking
- ✅ Reference implementations in Jupyter notebooks
| # | Dataset | Modality | Patients | #CT Scans | Condition | Demographics | Status |
|---|---|---|---|---|---|---|---|
| 1 | NSCLC-Radiomics (NSCLCR) | CT | 421 | 421 | Lung Cancer | Netherlands | ✅ Available |
| 2 | UniToChest | CT | 623 | 715 | Lung Nodules | Italy | ✅ Available |
| 3 | IMDCT | CT | 2,032 | 2,032 | Indeterminate Pulmonary Nodules | China (Multi-institutional) | ✅ Available |
| 4 | LNDb v4 | CT | 294 | 294 | Pulmonary Nodules | Portugal | ✅ Available |
| 5 | BIMCV-R | CT | 5,340 | 8,069 | Multi-label Findings (COVID-19, Pneumonia, etc.) | Spain | ✅ Available |
| 6 | LUNGx | CT | 83 | 83 | Lung Nodule Classification | USA | ✅ Available |
| 7 | LIDC-IDRI | CT | 875 | 875 | Lung Nodule Detection & Segmentation | USA (Multi-institutional) | ✅ Available |
| 8 | LUNA25 | CT | 2,020 | 4,069 | Lung Nodule Detection with AI Segmentations | Netherlands (Multi-institutional) | ✅ Available |
| 9 | MIDRC-RICORD | CT | 227 | 328 | COVID-19 Detection & Classification | USA (Multi-institutional) | ✅ Available |
| 10 | U-10 (United-10) | CT | 12,000+ | 12,000+ | COVID-19 Multi-Dataset Collection (10 datasets) | Multi-national | ✅ Available |
| 11 | NLST-3D | CT | 900+ | 969 | Lung Cancer Screening with 3D Nodule Annotations | USA (Multi-institutional) | ✅ Available |
| 12 | DLCS 2024 | CT (LDCT) | 2,061 | 2,061 | Lung Cancer Screening with Lung-RADS Scores | USA (Duke University) | ✅ Available |
| 13 | DeepLesion-1kTest3D | CT | 10,594 | 1,000 | Universal Lesion Detection (8 Body Regions) | USA (NIH Clinical Center) | ✅ Available |
| 14 | NLST ML Metadataset | Metadata | CT Screening Arm | Nodule-level | Lung Cancer Risk Prediction | USA (Multi-institutional) | ✅ Available |
| More datasets coming soon... | - | - | - | - | - | 🔜 Planned |
🎯 SEGMENTATION 🧠 CLASSIFICATION 🔍 DETECTION 📊 BENCHMARK 🧬 GENERATION ✍️ ANNOTATION
| 🔗 Resource | 🧠 Type | 📝 Details | 🌐 Link |
|---|---|---|---|
| VLST (Virtual Lung Screening Trials) | 🧬 Simulation | In-silico lung screening trials | https://fitushar.github.io/VLST.github.io/ |
| Lung Cancer Benchmarks | 📊 Classification, 🔍Detection | CADe & CADx Benchmarking | https://zenodo.org/records/13799069 · https://shorturl.at/Xh2uO |
| PiNS | 🎯 Segmentation | Point-driven nodule masks | https://github.com/fitushar/PiNS |
| CaNA | 🧩 Augmentation | Context-aware nodule synthesis | https://github.com/fitushar/CaNA |
| NoMAISI | 🧬 Syntheis | Controlled synthetic lung nodules | https://github.com/fitushar/NoMAISI |
| RBA | 🧠 NLP | Radiology text/report label extractor | https://github.com/fitushar/multi-label-weakly-supervised-classification-of-body-ct#rule-based-algorithm-rba |
| TriAnnot | 🧠 Annotation-tool | Tri-stage annotation consensus | 🚧 In preparation |
| 🔗 Resource | 👤 Contributor | 🧠 Type | 📝 Details | 🌐 Link |
|---|---|---|---|---|
| 3D-Segmentation Datasets | Lavsen Dahal | 🎯 Segmentation | Collection of open-source 3D segmentation models | https://github.com/lavsendahal/3D-segmentation |
| 3D Medical Imaging Preprocessing All you need | Fakrul Islam Tushar | 🎯 Pre-procising | Basic Tutorials | https://github.com/fitushar/3D-Medical-Imaging-Preprocessing-All-you-need |
| Automatic Breast Region Extractor | Fakrul Islam Tushar | 🎯 Pre-procising | Breast region extarctor using OpenCV | https://github.com/fitushar/Automatic_Breast_Region_Extraction_using_python/tree/master |
Non-Small Cell Lung Cancer CT Imaging with Radiomics Analysis
- Modality: CT (Computed Tomography)
- Patients: 421 (1 excluded due to segmentation issues)
- Condition: Non-Small Cell Lung Cancer (NSCLC)
- Source: TCIA - The Cancer Imaging Archive
- Original Publication: Aerts et al. (2015) - DOI: 10.7937/K9/TCIA.2015.PF0M9REI
- Pre-treatment CT scans: 512×512×[88-176] voxels, ~0.977mm in-plane, 3mm slice thickness
- Multi-label segmentations:
- Gross Tumor Volume (GTV-1)
- Esophagus, Heart, Spinal Cord
- Left & Right Lungs
- Clinical metadata: TNM staging, histology, survival time, demographics
- Bounding box annotations for tumor detection tasks
- Train/Validation/Test splits: 334/41/46 patients (stratified by histology)
- Preprocessed Dataset: Zenodo Repository - DOI: 10.5281/zenodo.18284366
- Processing Documentation: DATASET_PROCESSING_DOCUMENTATION.md
- Processing Notebook: NSCLCR_HAID_processing.ipynb
If you use this dataset, please cite:
@article{aerts2014decoding,
title={Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach},
author={Aerts, Hugo JWL and Velazquez, Emmanuel Rios and Leijenaar, Ralph TH and others},
journal={Nature Communications},
volume={5},
pages={4006},
year={2014},
doi={10.1038/ncomms5006}
}
@misc{aerts2015nsclc,
author={Aerts, Hugo JWL and Wee, Leonard and Rios Velazquez, Emmanuel and others},
title={Data From NSCLC-Radiomics},
year={2015},
publisher={The Cancer Imaging Archive},
doi={10.7937/K9/TCIA.2015.PF0M9REI}
}Chest CT Imaging with Expert-Annotated Lung Nodule Segmentation
- Modality: CT (Computed Tomography)
- Patients: 623 unique patients (715 CT scans total)
- Condition: Lung Nodules (10,071 annotated nodules)
- Source: Zenodo Repository
- Original Publication: Chaudhry et al. (2022) - ICIAP 2022 - DOI: 10.1007/978-3-031-06427-2_16
- Institution: Città della Salute e della Scienza di Torino & University of Turin, Italy
- Chest CT scans: 715 scans from 623 patients with varying slice thickness and spacing
- Expert annotations: 10,071 lung nodule segmentation masks by expert radiologists
- Standardized preprocessing: Resampled to uniform spacing [0.703125, 0.703125, 1.25] mm
- 3D bounding box annotations for nodule detection tasks
- Train/Validation/Test splits: 579/66/70 scans from 501/62/63 patients (~80%/10%/10%)
- Clinical metadata: Patient demographics (age, sex), scanner parameters, acquisition details
- Nodule characteristics: Diameter statistics (mean 21.4 ± 20.2 mm)
- Original Dataset: Zenodo Repository - DOI: 10.5281/zenodo.5797912
- Preprocessed Dataset: Zenodo Repository - DOI: 10.5281/zenodo.18285682
- Processing Documentation: UNITOCHEST_PROCESSING_DOCUMENTATION.md
- Processing Notebook: Demo_Dicom_to_CT-HAID.ipynb
- Data Analysis Notebook: DataAnalysis.ipynb
If you use this dataset, please cite:
@inproceedings{chaudhry2022unitochest,
title={UniToChest: A Lung Image Dataset for Segmentation of Cancerous Nodules on CT Scans},
author={Chaudhry, Aymen and Perlo, Daniele and Renzulli, Riccardo and Santinelli, Francesca and Tibaldi, Stefano and Cristiano, Carmen and Grosso, Marco and Limerutti, Giorgio and Grangetto, Marco and Fonio, Paolo},
booktitle={International Conference on Image Analysis and Processing (ICIAP)},
pages={189--200},
year={2022},
publisher={Springer},
doi={10.1007/978-3-031-06427-2_16}
}
@dataset{perlo2021unitochest,
author={Perlo, Daniele and Renzulli, Riccardo and Santinelli, Francesca and Tibaldi, Stefano and Cristiano, Carmen and Grosso, Marco and Limerutti, Giorgio and Grangetto, Marco and Fonio, Paolo},
title={UniToChest},
year={2021},
publisher={Zenodo},
doi={10.5281/zenodo.5797912}
}Indeterminate Pulmonary Nodules Multi-institutional CT Dataset with Histopathological Confirmation
- Modality: CT (Computed Tomography)
- Patients: 2,032 unique patients
- Condition: Indeterminate Pulmonary Nodules with histopathological confirmation
- Source: Zenodo Repository
- Original Publication: Zhao et al. (2024) - Nature Communications - DOI: 10.1038/s41467-024-55594-4
- Institutions: Multi-institutional (5 clinical centers in China)
- Chest CT scans: 2,032 scans with indeterminate pulmonary nodules
- Histopathological confirmation: 100% pathology-confirmed diagnoses (gold standard)
- One annotated nodule per patient: Single target per scan for simplified analysis
- High malignancy rate: 78.6% malignant, 21.4% benign nodules
- Clinically relevant sizes: Predominantly intermediate to very large nodules (10-30+ mm)
- 3D bounding box annotations for nodule detection and localization
- Train/Validation/Test splits: 1,625/203/204 patients (~80%/10%/10%)
- Clinical metadata: Patient age, nodule dimensions, malignancy labels
- Nodule characteristics: Mean diameter 30.0 ± 20.8 mm, solid component 15.7 ± 13.2 mm
- Original Dataset Part-I: Zenodo Repository - DOI: 10.5281/zenodo.13908015
- Original Dataset Part-II: Zenodo Repository - DOI: 10.5281/zenodo.13913777
- Processing Documentation: IMDCT_DATASET_DOCUMENTATION.md
- Processing Notebooks:
If you use this dataset, please cite:
@article{zhao2024integrated,
title={Integrated multiomics signatures to optimize the accurate diagnosis of lung cancer},
author={Zhao, Mengmeng and She, Yunlang and others},
journal={Nature Communications},
volume={15},
year={2024},
publisher={Nature Publishing Group},
doi={10.1038/s41467-024-55594-4}
}
@dataset{zhao2024imdct,
author={Zhao, Mengmeng and She, Yunlang},
title={CT dataset for "Integrated multiomics signatures to optimize the accurate diagnosis of lung cancer" Part-I},
year={2024},
publisher={Zenodo},
doi={10.5281/zenodo.13908015}
}Lung Nodule Database with Multi-Reader Expert Annotations and Text Report Extractions
- Modality: CT (Computed Tomography)
- Patients: 294 unique patients (237 after quality control)
- Condition: Pulmonary Nodules with multi-reader annotations
- Source: LNDb Grand Challenge
- Original Publication: Ferreira et al. (2024) - Scientific Data - DOI: 10.1038/s41597-024-03345-6
- Institutions: INESC TEC & São João Hospital Centre, Porto, Portugal
- Chest CT scans: 294 scans with comprehensive pulmonary nodule coverage
- Multi-reader annotations: Up to 4 expert radiologists per case (1,235 total nodule instances)
- Dual annotation sources: Radiological annotations + text report extractions
- Rich clinical attributes: 9 radiological characteristics rated on 1-6 scales:
- Texture, Calcification, Malignancy, Subtlety, Lobulation, Margin, Sphericity, Spiculation, Internal Structure
- Standardized preprocessing: Resampled to uniform spacing [0.703125, 0.703125, 1.25] mm
- Nodule size range: 3-40 mm diameter (~85% sub-centimeter)
- Train/Validation/Test splits: 189/24/24 patients (~80%/10%/10%)
- 3D bounding box annotations for nodule detection tasks
- Fleischner score compatibility: Suitable for patient follow-up recommendation systems
- Original Dataset: LNDb Grand Challenge Download
- Preprocessed Dataset: Available on Zenodo (DOI to be added)
- Processing Documentation: LNDBV4_DATASET_DOCUMENTATION.md
- Processing Notebook: LNDbv4-Datasetprocessing-HAID.ipynb
- Conversion Scripts:
- mhd_to_nifti.py - Convert CT scans
- mhd_to_nifti_masks.py - Convert segmentation masks
If you use this dataset, please cite:
@article{ferreira2024lndb,
title={LNDb v4: An open lung nodule database with multi-reader annotations and clinical attributes},
author={Ferreira, Carlos and Pedrosa, João and Aresta, Guilherme and others},
journal={Scientific Data},
volume={11},
year={2024},
publisher={Nature Publishing Group},
doi={10.1038/s41597-024-03345-6}
}
@misc{lndb_grandchallenge,
title={LNDb Challenge - Lung Nodule Database},
author={INESC TEC and São João Hospital Centre},
year={2020},
howpublished={\url{https://lndb.grand-challenge.org/}}
}Large-Scale Chest CT Dataset with Bilingual Radiology Reports and Multi-Label Findings
- Modality: CT (Computed Tomography)
- Patients: 5,340 unique patients (8,069 CT scans total)
- Condition: Multi-label radiological findings with comprehensive radiology reports
- Source: BIMCV Database | Hugging Face Dataset
- Original Publication: Chen et al. (2024) - MICCAI 2024 - DOI: 10.1007/978-3-031-72120-5_12
- Institution: Medical Imaging Databank of the Valencia Region (BIMCV), Spain
- Large-scale chest CT dataset: 8,069 CT volumes from 5,340 patients
- Bilingual radiology reports: Spanish (original) + English (machine translated)
- Rich multi-label annotations: 10+ major radiological findings including:
- Ground Glass Pattern (34.4%), Pneumonia (30.6%), COVID-19 (26.7%)
- Nodules (24.4%), Pleural Effusion (19.9%), Consolidation (12.5%)
- Adenopathy, Calcified Densities, Vertebral Changes, and more
- Real-world clinical data: Hospital data from Valencia, Spain
- COVID-19 cohort: Significant representation (2,152 instances, 26.7%)
- Text-image pairs: Over 2 million 2D CT slices with associated reports
- Proposed Train/Validation/Test splits: 4,272/534/534 patients (~80%/10%/10%)
- Patient-level stratification: No patient overlap across splits
- Original Dataset: Hugging Face - BIMCV-R
- BIMCV Database: BIMCV COVID-19 Projects
- Processing Documentation: BIMCVR_DATASET_DOCUMENTATION.md
- Analysis Notebook: BIMCV_DataAnalysis.ipynb
- Split Metadata: Available in metadata/ folder
If you use this dataset, please cite:
@inproceedings{chen2024bimcv,
title={BIMCV-R: A landmark dataset for 3D CT text-image retrieval},
author={Chen, Yinda and Liu, Che and Liu, Xiaoyu and Arcucci, Rossella and Xiong, Zhiwei},
booktitle={International Conference on Medical Image Computing and Computer-Assisted Intervention},
pages={124--134},
year={2024},
organization={Springer},
doi={10.1007/978-3-031-72120-5_12}
}
@misc{bimcv_database,
title={BIMCV-COVID19: Medical Imaging Databank of the Valencia Region},
author={BIMCV},
year={2020},
howpublished={\url{https://bimcv.cipf.es/bimcv-projects/bimcv-covid19/}}
}SPIE-AAPM-NCI Lung Nodule Classification Challenge Dataset with Pathology-Confirmed Diagnoses
- Modality: CT (Computed Tomography)
- Patients: 73 test patients + 10 calibration patients = 83 total
- Condition: Lung nodule classification (benign vs. malignant)
- Source: TCIA - The Cancer Imaging Archive
- Original Publication: Armato et al. (2016) - Journal of Medical Imaging - DOI: 10.1117/1.JMI.3.4.044506
- Challenge Organization: SPIE-AAPM-NCI (USA)
- DOI: 10.7937/K9/TCIA.2015.UZLSU3FL
- Diagnostic CT scans: 83 chest CT volumes with nodule annotations
- Pathology-confirmed labels: 100% histologically confirmed diagnoses (gold standard)
- Challenge dataset: Real-world diagnostic performance evaluation
- Nodule annotations: 94 nodules with precise DICOM coordinates
- Diagnosis distribution: 57 benign (60.6%), 37 malignant (39.4%)
- Multi-nodule cases: 11 patients with 2 nodules each
- Scanner: Philips CT scanners (high-resolution protocols)
- Processed features:
- World coordinate conversion from DICOM pixel space
- Standardized resampling to [0.703125, 0.703125, 1.25] mm
- 64×64×64 voxel diagnostic patches centered on nodules
- Original Dataset: TCIA - SPIE-AAPM Lung CT Challenge
- Nodule Annotations:
- Processing Documentation: LUNGX_DATASET_DOCUMENTATION.md
- Processing Notebook: LUNGx-Processing-HAID.ipynb
If you use this dataset, please cite:
@article{armato2016lungx,
title={LUNGx Challenge for computerized lung nodule classification},
author={Armato III, Samuel G and Drukker, Karen and Li, Feng and Hadjiiski, Lubomir and Tourassi, Georgia D and Engelmann, Roger M and Giger, Maryellen L and Redmond, George and Farahani, Keyvan and Kirby, Justin S and Clarke, Laurence P},
journal={Journal of Medical Imaging},
volume={3},
number={4},
pages={044506},
year={2016},
publisher={SPIE},
doi={10.1117/1.JMI.3.4.044506}
}
@dataset{armato2015lungx,
author={Armato III, Samuel G and Hadjiiski, Lubomir and Tourassi, Georgia D and Drukker, Karen and Giger, Maryellen L and Li, Feng and Redmond, George and Farahani, Keyvan and Kirby, Justin S and Clarke, Laurence P},
title={SPIE-AAPM-NCI Lung Nodule Classification Challenge Dataset},
year={2015},
publisher={The Cancer Imaging Archive},
doi={10.7937/K9/TCIA.2015.UZLSU3FL}
}Lung Image Database Consortium - Image Database Resource Initiative with Multi-Radiologist Consensus Annotations
- Modality: CT (Computed Tomography)
- Patients: 875 unique patients (processed from original 1,010)
- Condition: Lung nodule detection and segmentation
- Source: TCIA - The Cancer Imaging Archive
- Original Publication: Armato et al. (2011) - Medical Physics - DOI: 10.1118/1.3528204
- Consortium: LIDC-IDRI (7 academic centers, NCI/NIH/FDA partnership, USA)
- DOI: 10.7937/K9/TCIA.2015.LO9QL9SX
- Comprehensive lung nodule benchmark: 875 diagnostic and screening chest CT scans
- Multi-radiologist annotations: Up to 4 expert thoracic radiologists per case
- Two-phase annotation process: Blinded read followed by unblinded review
- Consensus approach: Union of all radiologist annotations for complete nodule coverage
- Rich nodule characterization: Size, texture, calcification, malignancy, spiculation, lobulation ratings
- Multi-class segmentation: Nodule + organ (Vista3D) + body mask integration
- Dataset splits: 649 train / 163 validation / 63 test patients
- Label scheme:
- Label 23: Pulmonary nodules (multi-radiologist union)
- Label 200: Body/soft tissue regions
- Other labels: Vista3D organ segmentations
- Standardized format: NIfTI with comprehensive metadata
- Original Dataset: TCIA - LIDC-IDRI Collection
- Radiologist Annotations: XML Files (8.62MB)
- pylidc Tool: GitHub - pylidc (recommended for working with annotations)
- Processing Documentation: LIDCIDRI_DATASET_DOCUMENTATION.md
- Processing Notebook: LIDC-IDRI-Segmentation-Processing-HAID.ipynb
If you use this dataset, please cite:
@article{armato2011lidc,
title={The Lung Image Database Consortium (LIDC) and Image Database Resource Initiative (IDRI): a completed reference database of lung nodules on CT scans},
author={Armato III, Samuel G and McLennan, Geoffrey and Bidaut, Luc and McNitt-Gray, Michael F and Meyer, Charles R and Reeves, Anthony P and Zhao, Binsheng and Aberle, Denise R and Henschke, Claudia I and Hoffman, Eric A and others},
journal={Medical Physics},
volume={38},
number={2},
pages={915--931},
year={2011},
publisher={Wiley Online Library},
doi={10.1118/1.3528204}
}
@dataset{armato2015lidc,
author={Armato III, Samuel G and McLennan, Geoffrey and Bidaut, Luc and McNitt-Gray, Michael F and Meyer, Charles R and Reeves, Anthony P and others},
title={Data From LIDC-IDRI},
year={2015},
publisher={The Cancer Imaging Archive},
doi={10.7937/K9/TCIA.2015.LO9QL9SX}
}Acknowledgement: The authors acknowledge the National Cancer Institute and the Foundation for the National Institutes of Health, and their critical role in the creation of the free publicly available LIDC/IDRI Database used in this study.
Extended LUNA16 with Deep Learning-Based Nodule and Organ Segmentation
- Modality: CT (Computed Tomography)
- Patients: 2,020 unique patients (4,069 CT scans)
- Condition: Lung nodule detection with comprehensive AI-generated segmentations
- Source: LUNA16 Grand Challenge
- Original Publication: Setio et al. (2017) - Medical Image Analysis - DOI: 10.1016/j.media.2017.06.015
- Organizers: Radboud University Medical Center, Nijmegen, Netherlands
- Segmentation Models: PiNS + MONAI Vista3D
- Extended LUNA16 dataset: LUNA25 = LUNA16 + Deep Learning Segmentations
- 6,163 nodule annotations: Complete nodule set with world coordinates (x, y, z in mm)
- AI-powered nodule segmentation: PiNS deep learning model for precise nodule boundaries
- Comprehensive organ segmentation: MONAI Vista3D foundation model for multi-organ context
- Integrated multi-class masks: Nodule + organ + body segmentation in single NIfTI files
- Dataset splits: 4,930 train / 617 validation / 616 test nodule annotations
- Label scheme:
- Label 23: Pulmonary nodules (PiNS segmentation with morphological erosion)
- Label 200: Body/soft tissue regions
- Other labels: Vista3D organ segmentations (lungs, airways, vessels, heart, etc.)
- Morphological refinement: 3×3×3 erosion kernel for high-confidence nodule cores
- KNN expansion option: 2mm K-nearest-neighbor dilation for nodule margin inclusion
- Clinical metadata: Age, gender, study dates, nodule coordinates
- PyRadiomics features: Optional radiomics feature extraction for nodule characterization
- Original LUNA16: Grand Challenge - LUNA16
- Preprocessed Segmentations: Zenodo - DOI: 10.5281/zenodo.18291811
- PiNS Segmentation Model: GitHub - PiNS
- MONAI Vista3D: GitHub - MONAI VISTA
- Processing Documentation: LUNA25_DATASET_DOCUMENTATION.md
- Processing Notebooks:
If you use this dataset, please cite:
@article{setio2017validation,
title={Validation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: The LUNA16 challenge},
author={Setio, Arnaud Arindra Adiyoso and Traverso, Alberto and De Bel, Thomas and others},
journal={Medical Image Analysis},
volume={42},
pages={1--13},
year={2017},
publisher={Elsevier},
doi={10.1016/j.media.2017.06.015}
}
@misc{tushar2026luna25,
author={Fakrul Islam Tushar},
title={LUNA25: Extended LUNA16 with Deep Learning Segmentations},
year={2026},
publisher={Zenodo},
doi={10.5281/zenodo.18291811},
note={PiNS + MONAI Vista3D Segmentations}
}
@misc{tushar2024pins,
author={Fakrul Islam Tushar},
title={PiNS: Precise Nodule Segmentation for Lung Cancer Detection},
year={2024},
publisher={GitHub},
howpublished={\url{https://github.com/fitushar/PiNS}}
}Key Innovation: LUNA25 extends the widely-used LUNA16 benchmark with state-of-the-art deep learning segmentations (PiNS for nodules, Vista3D for organs), providing researchers with precise anatomical context for AI model development.
Multi-Institutional COVID-19 Chest CT Database with RT-PCR Confirmation
- Modality: CT (Computed Tomography)
- Patients: 227 unique patients (328 CT scans)
- Condition: COVID-19 detection and classification
- Source: MIDRC - Medical Imaging Data Resource Center
- Dataset Versions: MIDRC-RICORD-1A (COVID-19 positive) + MIDRC-RICORD-1B (COVID-19 negative controls)
- Consortium: RSNA International COVID-19 Open Radiology Database (RICORD), USA
- Data Collection Period: 2020-2021 (COVID-19 pandemic)
- MIDRC-RICORD-1A: COVID-19 positive cases confirmed by RT-PCR
- MIDRC-RICORD-1B: COVID-19 negative control cases
- Binary classification: COVID-19 detected vs. not detected
- Clinical metadata: Age, gender, study dates, specimen source (pooled NP/OP swab)
- RT-PCR confirmation: Gold standard COVID-19 testing for positive cases
- Multi-institutional: Data from multiple US medical centers
- Dataset splits: 196 train / 57 validation / 75 test CT scans
- Standardized preprocessing: DICOM to NIfTI conversion with consistent formatting
- Variable slice thickness: 1.25mm - 3.0mm (clinical acquisition protocols)
- Image orientations: Axial (original), Coronal/Sagittal (reformatted views available)
- Symptomatic status: Available for subset of patients in RICORD-1B
- Original MIDRC Portal: https://www.midrc.org/
- RICORD Information: Multi-institutional contribution from RSNA collaboration
- Processing Scripts:
- Metadata Files:
If you use this dataset, please cite:
@misc{midrc2021ricord,
author={Medical Imaging Data Resource Center (MIDRC)},
title={RSNA International COVID-19 Open Radiology Database (RICORD)},
year={2021},
publisher={MIDRC},
howpublished={\url{https://www.midrc.org/}},
note={Multi-institutional COVID-19 chest CT database}
}
@article{tsai2021rsna,
title={The RSNA International COVID-19 Open Radiology Database (RICORD)},
author={Tsai, Eduardo Baptista and Simpson, Sheela and Lungren, Matthew P and others},
journal={Radiology},
volume={299},
number={1},
pages={E204--E213},
year={2021},
publisher={Radiological Society of North America},
doi={10.1148/radiol.2021203957}
}Clinical Significance: MIDRC-RICORD provides RT-PCR confirmed COVID-19 cases with matched negative controls, enabling development and validation of AI models for COVID-19 detection and severity assessment from chest CT imaging.
Unified Collection of 10 Public COVID-19 CT Datasets with Virtual Imaging Trials
- Modality: CT (Computed Tomography)
- Total Volumes: 12,000+ CT volumes from 10 public datasets
- Condition: COVID-19 detection and diagnosis with Virtual Imaging Trials (VIT)
- Source: Zenodo Repository - DOI: 10.5281/zenodo.14064172
- Original Publication: Tushar et al. (2023) - arXiv:2308.09730
- Processed By: Fakrul Islam Tushar (Duke University)
- Project Page: https://fitushar.github.io/ReviCOVID.github.io/
- Unified multi-dataset collection: Aggregates 10 publicly available COVID-19 CT datasets
- 12,000+ CT volumes: Comprehensive coverage across diverse populations and protocols
- 10 source datasets:
- BIMCV (Spain)
- COVIDx CT-2A (Multi-national)
- COVID-CT-MD (Multi-national)
- CT-NIH (USA)
- Lung Effusion/NSCLC (Multi-purpose)
- Lung Cancer (Various)
- LIDC-IDRI (USA)
- MIDRC-RICORD (USA)
- MosMed (Russia)
- NY Subset (USA)
- Virtual Imaging Trials (VIT): Includes simulated CT data using XCAT phantoms and DukeSim framework
- Pre-processed TFRecords: Ready-to-use TensorFlow format (96×160×160 voxels)
- Train/Validation/Test splits: Pre-defined splits for reproducible research
- Diverse imaging protocols: Multiple scanner configurations, acquisition parameters
- Multi-institutional: Data from hospitals and research centers across multiple countries
- COVID-19 labeling: Binary classification (COVID-19 positive/negative)
- Zenodo Repository: DOI: 10.5281/zenodo.14064172
- Total Size: 52.5 GB (10 zip files with pre-processed TFRecords)
- Project Page: ReviCOVID Project
- Source Code:
- Virtual Imaging Data: Available through CVIT Portal
- bimcv_tfrecords_96x160x160.zip (24.6 GB)
- covidctdata_tfrecords_96x160x160.zip (3.7 GB)
- covidctmd_tfrecords_96x160x160.zip (2.7 GB)
- ct_NIHa_tfrecords_96x160x160.zip (2.3 GB)
- effusion_nclcl_tfrecords_96x160x160.zip (1.8 GB)
- lgcancer_tfrecords_96x160x160.zip (2.9 GB)
- lidi_idri_tfrecords_96x160x160.zip (3.9 GB)
- midric_ricord_tfrecords_96x160x160.zip (1.0 GB)
- mosmed_tfrecords_96x160x160.zip (4.5 GB)
- ny_sub_tfrecords_96x160x160.zip (5.0 GB)
If you use this dataset, please cite:
@misc{tushar2024u10,
author={Fakrul Islam Tushar},
title={U-10: United-10 COVID19 CT Dataset},
year={2024},
publisher={Zenodo},
doi={10.5281/zenodo.14064172},
note={Virtual imaging trials improved the transparency and reliability of AI systems in COVID-19 imaging}
}
@article{tushar2023virtual,
title={Virtual Imaging Trials Improved the Transparency and Reliability of AI Systems in COVID-19 Imaging},
author={Tushar, Fakrul Islam and Dahal, Lavsen and Sotoudeh-Paima, Saman and Abadi, Ehsan and Segars, W. Paul and Samei, Ehsan and Lo, Joseph Y.},
journal={arXiv preprint arXiv:2308.09730},
year={2023}
}
@inproceedings{tushar2022virtual,
title={Virtual vs. reality: external validation of COVID-19 classifiers using XCAT phantoms for chest computed tomography},
author={Tushar, Fakrul Islam and Abadi, Ehsan and Sotoudeh-Paima, Saman and Fricks, Rafael B and Mazurowski, Maciej A and Segars, W Paul and Samei, Ehsan and Lo, Joseph Y},
booktitle={Medical Imaging 2022: Computer-Aided Diagnosis},
volume={12033},
pages={1203305},
year={2022},
organization={SPIE},
doi={10.1117/12.2613010}
}Research Impact: U-10 enables large-scale COVID-19 AI research by unifying 10 diverse datasets with standardized preprocessing. The integration of Virtual Imaging Trials provides controlled experiments to evaluate AI model transparency, reliability, and generalizability across different imaging conditions.
National Lung Screening Trial with 3D Nodule Annotations
- Modality: CT (Computed Tomography)
- Patients: 900+ lung cancer patients
- CT Scans: 969 annotated CT scans
- Condition: Lung cancer screening with 3D nodule detection
- Source: National Lung Screening Trial (NLST)
- Original Study: National Lung Screening Trial Research Team (2011) - NEJM
- Original 2D Annotations: Mikhael et al. (2023) - DOI: 10.1200/JCO.22.01345
- NLST-3D Annotations: DOI: 10.5281/zenodo.15320923
- Details & Visualization: GitHub - NLST Data Annotations
- 3D Conversion: Fakrul Islam Tushar (Duke University)
- NLST Background: Landmark randomized trial (53,454 participants from 33 US screening centers)
- 20% mortality reduction: Three annual low-dose CT screenings vs. radiography
- 969 annotated CT scans: From 900+ lung cancer patients
- 1,192 3D nodule annotations: Converted from 9,000+ 2D slice-level bounding boxes
- Annotation processing:
- Verified 2D annotations within DICOM images
- Extracted seriesinstanceuid, slice_location, slice_number from DICOM headers
- Converted image coordinates to world coordinates
- Verified annotations in corresponding NIfTI images
- Merged overlapping consecutive 2D annotations into single 3D annotations
- Multi-institutional data: 33 screening centers across the USA
- Clinical context: High-risk screening population (age 55-74, 30+ pack-year smoking history)
- Stage distribution: Emphasis on early-stage detection (47.5% stage IA in CT group)
- 3D Annotations (Zenodo): DOI: 10.5281/zenodo.15320923
- File:
fitetal_NLST_3D_Annotation_worldxyzwh_CenterCordxyz.csv(155.1 kB) - Columns: World coordinates (x, y, z), width, height, depth, center coordinates
- File:
- Original NLST Data: Available through Cancer Data Access System (CDAS)
- GitHub Repository: AI in Lung Health Benchmarking
- GitLab Repository: CVIT AI Lung Health Benchmarking
- Visualization Notebook: 3D Annotation Visualizations
If you use this dataset, please cite:
@dataset{tushar2025nlst3d,
author={Fakrul Islam Tushar},
title={NLST-3D+ Annotation},
year={2025},
publisher={Zenodo},
doi={10.5281/zenodo.15320923}
}
@article{mikhael2023ai,
title={AI-Based Lung Cancer Risk Prediction From National Lung Screening Trial Data},
author={Mikhael, Peter G and Wohlwend, Jeremy and Yala, Adam and others},
journal={Journal of Clinical Oncology},
volume={41},
number={23},
pages={3993--4004},
year={2023},
doi={10.1200/JCO.22.01345}
}
@article{nlst2011reduced,
title={Reduced Lung-Cancer Mortality with Low-Dose Computed Tomographic Screening},
author={{National Lung Screening Trial Research Team}},
journal={New England Journal of Medicine},
volume={365},
number={5},
pages={395--409},
year={2011},
doi={10.1056/NEJMoa1102873}
}
@article{aberle2013results,
title={Results of the Two Incidence Screenings in the National Lung Screening Trial},
author={Aberle, Denise R and DeMello, Sarah and Berg, Christine D and others},
journal={New England Journal of Medicine},
volume={369},
number={10},
pages={920--931},
year={2013},
doi={10.1056/NEJMoa1208962}
}Landmark Study: NLST demonstrated that annual low-dose CT screening reduces lung cancer mortality by 20% in high-risk populations. This 3D annotation dataset enables development of advanced nodule detection and characterization AI models using proven clinically significant screening data.
Duke Lung Cancer Screening Dataset with Contemporary CT Technology
- Modality: Low-Dose CT (LDCT)
- Total Patients: 2,061 patients from Duke University Health System
- Publicly Available: 1,613 CT scans with 2,487 annotated nodules
- Reserved Test Set: 448 scans (21.7%) withheld for future challenges
- Condition: Lung cancer screening with Lung-RADS classification
- Time Period: January 1, 2015 to June 30, 2021
- Source: RSNA Radiology: Artificial Intelligence
- Dataset Repository: Zenodo - DOI: 10.5281/zenodo.13799069
- Original Publication: Wang et al. (2025) - DOI: 10.1148/ryai.240248
- arXiv Preprint: 2405.04605
- Contemporary CT technology: First large dataset reflecting current (2015-2021) CT scanner technology and clinical practice
- Semiautomated annotations: 85.8% of scans (1,768 of 2,061) annotated semiautomatically
- Group 1: Fully automatic (33/34 = 97% accuracy)
- Group 2: Automatic with manual selection (32/33 = 97% accuracy)
- Group 3: Algorithm-detected, manually confirmed (45/45 = 100% accuracy)
- Group 4: Manual annotation with radiologist review (180/183 = 98% accuracy)
- Efficient annotation method: MONAI nodule detection algorithm trained on LUNA16 dataset
- Cross-validation sensitivity: 0.835 at 1/8 FP per scan, 0.988 at 8 FP per scan
- Reduced radiologist annotation time by >90%
- 3D bounding box annotations: Object (.obj) files with 8 coordinates per nodule
- Clinical scoring: Lung-RADS (Lung CT Screening Reporting and Data System) scores
- Versions 1.0 and 1.1 used during study period
- All potentially actionable nodules ≥4mm annotated
- Includes Lung-RADS 3, 4A, 4B, 4X categories
- Patient demographics: Mean age 66.7 ± 6.2 years (1,032 female, 1,029 male)
- Cancer outcomes: 155 patients (7.5%) with subsequent lung cancer diagnosis
- Includes timing, histologic type, and stage information
- Image format: NIfTI files with 0.6mm axial section thickness
- High-volume screening center: Data from Duke University's contemporary lung cancer screening program
- Reproducible methodology: Annotation pipeline can be replicated for future dataset creation
Note: Dataset access requires approval through Zenodo request form
- Part 1 (Primary Data): Zenodo - Subsets 1-7 + Metadata
- LDCT images (NIfTI format)
- Nodule annotations (.obj files)
- Metadata spreadsheet
- Part 2: Zenodo - Subsets 8-9
- Part 3: Zenodo - Subset 10
- RSNA Publication: Full Article
- arXiv Preprint: 2405.04605
- GitHub Repository: Available (see Zenodo record for links)
- GitLab Repository: Available (see Zenodo record for links)
If you use this dataset, please cite:
@article{wang2025dlcs,
title={The Duke Lung Cancer Screening (DLCS) Dataset: A Reference Dataset of Annotated Low-Dose Screening Thoracic CT},
author={Wang, Avivah J and Tushar, Fakrul Islam and Harowicz, Michael R and Tong, Betty C and Lafata, Kyle J and Tailor, Tina D and Lo, Joseph Y},
journal={Radiology: Artificial Intelligence},
volume={7},
number={4},
pages={e240248},
year={2025},
publisher={Radiological Society of North America},
doi={10.1148/ryai.240248}
}
@dataset{wang2024dlcs_zenodo,
author={Wang, Avivah and Tushar, Fakrul Islam and Harowicz, Michael R and Lafata, Kyle J and Tailor, Tina D and Lo, Joseph Y},
title={Duke Lung Cancer Screening Dataset},
year={2024},
publisher={Zenodo},
version={1.1},
doi={10.5281/zenodo.13799069}
}
@misc{wang2024dlcs_arxiv,
title={The Duke Lung Cancer Screening Dataset: A Reference Dataset of Annotated Low-Dose Screening Thoracic CT},
author={Wang, Avivah J and Tushar, Fakrul Islam and Harowicz, Michael R and Tong, Betty C and Lafata, Kyle J and Tailor, Tina D and Lo, Joseph Y},
year={2024},
eprint={2405.04605},
archivePrefix={arXiv},
primaryClass={eess.IV}
}Funding Acknowledgment: This work was supported by the Duke Radiology Putman Vision Award, NIH/NIBIB P41-EB028744, and NIH/NCI R01-CA261457.
Key Innovation: DLCS 2024 bridges the gap between older datasets (LIDC-IDRI from 2000s) and contemporary clinical practice, providing the first large-scale lung cancer screening dataset that reflects current CT technology. The efficient semiautomated annotation methodology enables scalable dataset creation and reduces radiologist workload while maintaining high accuracy (>90%).
Universal Lesion Detection Dataset with Multi-Organ 3D Annotations
- Modality: CT (Computed Tomography)
- Original Dataset: 32,735 lesions from 32,120 CT scans (10,594 patients)
- Test Set (3D NIfTI): 1,000 CT volumes with 4,927 lesion annotations
- Condition: Universal lesion detection across 8 anatomical regions
- Source: NIH Clinical Center - DeepLesion
- 3D NIfTI Repository: Zenodo - DOI: 10.5281/zenodo.18292965
- Original Publication: Yan et al. (2018) - DOI: 10.1109/JBHI.2017.2780066
- 3D Conversion: Fakrul Islam Tushar (Duke University)
- Large-scale lesion database: Originally 32,735 diverse lesions from real-world clinical practice
- Test subset: 1,000 3D CT volumes with 4,927 annotated lesions
- Multi-lesion volumes: Many volumes contain 2-10+ lesions per scan
- 8 Coarse Lesion Categories:
- Type 1: Lung nodules (~1,500 lesions, 30%)
- Type 2: Abdomen lesions (~400 lesions, 8%)
- Type 3: Mediastinum/Lymph nodes (~800 lesions, 16%)
- Type 4: Liver lesions (701 lesions, 14%)
- Type 5: Soft tissue lesions (~500 lesions, 10%)
- Type 6: Kidney/urinary lesions (235 lesions, 5%)
- Type 7: Bone lesions (~600 lesions, 12%)
- Type 8: Pelvic lesions (~191 lesions, 4%)
- RECIST Measurements: Long and short axis diameters for each lesion
- Annotation Source: PACS system bookmarks from clinical radiology practice at NIH Clinical Center
- 3D Bounding Boxes: Precise lesion localization with normalized coordinates (0-1 range)
- Clinical Context: Real-world lesion annotations from routine clinical workflow
- Organ-Specific Subsets Available:
- Liver Lesions: 701 annotations (test set)
- Kidney Lesions: 235 annotations (test set)
- 16-bit Intensity Correction: Proper Hounsfield Unit restoration from PNG encoding
- Slice Context: Variable slice ranges (30-60 slices per volume, some up to 270 slices)
- Image Resolution: 512×512 pixels, in-plane spacing 0.31-0.98 mm/pixel, slice thickness 1.0-5.0mm
- Demographics: Age 11-87 years, mixed gender, diverse adult and pediatric population
- Quality Flags: ~95% clean annotations, ~5% potentially noisy
- 3D NIfTI Test Set (Zenodo): DOI: 10.5281/zenodo.18292965
- 1,000 3D CT volumes (NIfTI format)
- Test set metadata with lesion annotations
- Organ-specific subsets (liver, kidney)
- Original DeepLesion (NIH): DeepLesion Box Repository
- Full dataset: 32,735 lesions, 32,120 CT scans
- 16-bit PNG format with metadata
- Processing Documentation: DEEPLESION_DATASET_DOCUMENTATION.md
- Conversion Script: DL_save_nifti.py
- GitHub Repository: HAID - DeepLesion Processing
If you use this dataset, please cite:
@article{yan2018deeplesion,
title={DeepLesion: automated mining of large-scale lesion annotations and universal lesion detection with deep learning},
author={Yan, Ke and Wang, Xiaosong and Lu, Le and Summers, Ronald M},
journal={Journal of Biomedical and Health Informatics},
volume={22},
number={4},
pages={1091--1101},
year={2018},
publisher={IEEE},
doi={10.1109/JBHI.2017.2780066}
}
@dataset{tushar2026deeplesion3d,
author={Fakrul Islam Tushar},
title={DeepLesion-1kTest3D: 3D NIfTI Conversion of DeepLesion Test Set},
year={2026},
publisher={Zenodo},
doi={10.5281/zenodo.18292965},
url={https://doi.org/10.5281/zenodo.18292965}
}
@misc{yan2018deeplesion_nih,
author={Yan, Ke and Wang, Xiaosong and Lu, Le and Summers, Ronald M},
title={DeepLesion: Large-scale Lesion Annotations from CT with Deep Learning},
year={2018},
howpublished={NIH Clinical Center},
url={https://nihcc.app.box.com/v/DeepLesion}
}Unique Contribution: DeepLesion is the first large-scale universal lesion detection dataset spanning multiple body regions, enabling development of comprehensive lesion detection AI models. The 3D NIfTI conversion (DeepLesion-1kTest3D) facilitates volumetric analysis and integration with modern medical imaging pipelines. Real-world PACS bookmarks ensure clinical relevance, with lesions representing actual findings tracked by radiologists in routine practice.
National Lung Screening Trial - Machine Learning Ready Nodule Dataset
- Modality: Metadata (derived from CT scans)
- Participants: CT screening arm participants from NLST
- Record Type: Nodule-level analysis dataset
- Focus: Non-calcified nodules ≥ 4 mm (abnormality code 51)
- Clinical Task: Lung cancer risk prediction and nodule characterization
- Source: National Lung Screening Trial (NLST) - CDAS
- Original Study: National Lung Screening Trial Research Team (2011) - NEJM
- Processed By: Fakrul Islam Tushar (Duke University)
- Repository: GitHub - HAID
- Nodule-level granularity: Each row represents a single CT-detected nodule with complete clinical context
- Comprehensive metadata integration: Demographics, nodule imaging features, cancer outcomes, risk factors
- Machine learning ready: Cleaned, decoded, and engineered features for immediate ML deployment
- Temporal validation splits: Separated by NLST CT Set (Set-1 vs. Set-2) for robust model validation
- Inclusion criteria: Non-calcified nodules ≥ 4 mm with complete metadata (no missing values)
- Feature engineering: 8 clinically-relevant derived features including:
- Family history aggregation
- Nodule type categorization (solid/part-solid/ground-glass)
- Spiculation binary indicator
- Upper lobe location flag
- Cancer outcome labels
- Time-indexed cancer diagnosis flags
- Clinical risk factors: Age, smoking history, emphysema diagnosis, family history
- Nodule characteristics: Size (longest diameter), attenuation type, margin morphology, anatomical location
- Outcome tracking: Cancer screening results, cancer timing, primary cancer locations
Core Clinical Variables:
pid: Participant ID (unique patient identifier)study_yr: Screening year (0, 1, 2)age: Participant age at screeninggender: Male / Femalerace: Decoded race/ethnicityall_sct_set: CT Set assignment (1 or 2)diagemph: Emphysema diagnosis (0/1)Family_History: Family history of lung cancer (0/1)can_scr: Cancer screening outcome (No Cancer / Lung Cancer - Screening Detected / Clinical Detection / Other Cancer)
Nodule Imaging Features:
sct_long_dia: Nodule longest diameter (mm)sct_margins: Nodule margin characteristics (Smooth/Lobulated/Spiculated/Irregular)sct_pre_att: Nodule attenuation type (Solid/Part-solid/Ground-glass)sct_epi_loc: Anatomical lobe location (RUL/RML/RLL/LUL/LLL/Multiple)
Engineered Features:
Nodule_Type: Categorized nodule type (solid/part-solid/ground-glass)Spiculation: Binary spiculation indicator (0/1) - key malignancy predictorUpper_Lobe: Binary upper lobe location (0/1) - higher malignancy riskCancer_lbl: Binary cancer outcome (0 = No Cancer, 1 = Cancer)primary_cancer_locations: Comma-separated lung cancer locationslung_cancer_t0,lung_cancer_t1,lung_cancer_t2: Time-indexed cancer flagsmatch_primary: Does nodule location match diagnosed cancer location (0/1)
1. Data Extraction & Merging:
- Merged participant metadata with CT abnormality records
- Combined demographics, clinical history, and nodule detections
- Preserved nodule-level granularity for ML applications
2. Categorical Decoding:
- Converted numerical codes to human-readable labels
- Gender, race, screening group, cancer outcomes
- Nodule location, attenuation, margin characteristics
3. Cohort Filtering:
- CT screening arm only (
rndgroup == 1) - Non-calcified nodules ≥ 4mm (
sct_ab_desc == 51) - Complete metadata (no missing critical variables)
- Valid nodule types (excluded "others/unknown")
4. Feature Engineering:
- Family history aggregation across all relatives
- Nodule type standardization
- Spiculation extraction (key malignancy risk factor)
- Upper lobe binary flag (anatomical risk stratification)
- Cancer outcome labels for supervised learning
- Time-indexed cancer diagnosis flags
5. Dataset Splitting:
- Set-1: NLST CT Set-1 participants (temporal cohort 1)
- Set-2: NLST CT Set-2 participants (temporal cohort 2)
- Enables temporal validation and generalization assessment
- Processing Notebook: NLST_ML_Metadataset_prep.ipynb
- Documentation: NLST_ML_METADATASET_DOCUMENTATION.md
- Output Files (in
output_dir/):nlst_ct_nodule_df_set1.csv: Set-1 nodule-level datasetnlst_ct_nodule_df_set2.csv: Set-2 nodule-level dataset
- Source Data: Requires approved access through CDAS (Cancer Data Access System)
- Input Files (in
scr_dir/):participant_d040722.csv: Participant demographics and outcomesSpiral CT Abnormalities/sct_abnormalities_d040722.csv: CT nodule detections
1. Lung Cancer Risk Prediction:
- Train ML models to predict malignancy risk from nodule + clinical features
- Binary classification: Cancer vs. No Cancer
- Feature importance analysis for clinical decision support
2. Nodule Characterization Analysis:
- Compare solid vs. subsolid nodule cancer risk profiles
- Evaluate spiculation as predictive biomarker
- Assess location-based risk stratification (upper lobe effect)
3. Temporal Validation:
- Train on Set-1, validate on Set-2 (or vice versa)
- Assess model generalization across temporal cohorts
- Detect temporal biases or population shifts
4. Risk Stratification Models:
- Combine clinical factors (age, smoking, family history, emphysema)
- Integrate nodule imaging features (size, type, location, margins)
- Develop composite risk scores for clinical translation
5. Feature Engineering Research:
- Identify strongest predictors of malignancy
- Compare demographic vs. imaging feature contributions
- Guide development of parsimonious clinical decision tools
If you use this dataset, please cite:
@misc{tushar2026nlst_ml_metadataset,
author={Fakrul Islam Tushar},
title={NLST ML Metadataset: Machine Learning Ready Nodule-Level Dataset from National Lung Screening Trial},
year={2026},
publisher={GitHub},
howpublished={\url{https://github.com/fitushar/HAID}}
}
@article{nlst2011reduced,
title={Reduced Lung-Cancer Mortality with Low-Dose Computed Tomographic Screening},
author={{National Lung Screening Trial Research Team}},
journal={New England Journal of Medicine},
volume={365},
number={5},
pages={395--409},
year={2011},
doi={10.1056/NEJMoa1102873}
}
@article{aberle2013results,
title={Results of the Two Incidence Screenings in the National Lung Screening Trial},
author={Aberle, Denise R and DeMello, Sarah and Berg, Christine D and others},
journal={New England Journal of Medicine},
volume={369},
number={10},
pages={920--931},
year={2013},
doi={10.1056/NEJMoa1208962}
}Clinical Context: NLST demonstrated a landmark 20% reduction in lung cancer mortality through CT screening in high-risk populations. This metadataset enables researchers to develop machine learning models for lung cancer risk prediction using validated clinical data, with robust features for nodule characterization and outcome prediction.
Key Innovation: First nodule-level NLST dataset with comprehensive feature engineering and temporal validation splits, bridging clinical metadata with modern ML pipelines for lung cancer risk assessment.
- Browse available datasets in the table above
- Download preprocessed data from the provided Zenodo links
- Review processing documentation in each dataset folder
- Load data splits using the provided CSV/JSON files
- Run reference notebooks to understand preprocessing pipelines
We welcome contributions! If you'd like to add a new dataset or improve existing documentation:
- Fork this repository
- Create a feature branch (
git checkout -b feature/new-dataset) - Follow our contribution guidelines
- Submit a pull request
This repository and preprocessing code are licensed under the Apache License 2.0.
Individual datasets retain their original licenses:
- NSCLC-Radiomics: Creative Commons Attribution 3.0 Unported License
- UniToChest: Creative Commons Attribution 4.0 International License
- IMDCT: Creative Commons Attribution 4.0 International License
- LNDb v4: Creative Commons Attribution 4.0 International License
- BIMCV-R: MIT License (academic research purposes only)
- LUNGx: Creative Commons Attribution 3.0 Unported License
- LIDC-IDRI: Creative Commons Attribution 3.0 Unported License
- LUNA25: Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License
- MIDRC-RICORD: Creative Commons Attribution 4.0 International License
- U-10 (United-10): Creative Commons Attribution 4.0 International License
- NLST-3D: Creative Commons Attribution 4.0 International License
- DLCS 2024: Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License
- DeepLesion-1kTest3D: Creative Commons Attribution 4.0 International License
Please review each dataset's specific license before use.
Maintainer: Fakrul Islam Tushar, PhD
Email: fitushar.mi@gmail.com
GitHub: @fitushar
For dataset-specific questions, please open an issue in this repository.
- TCIA (The Cancer Imaging Archive) for hosting and maintaining public medical imaging datasets
- Original dataset authors and contributors
- Open-source medical imaging community
- v1.1.0 (February 6, 2026): Added NLST ML Metadataset
- NLST ML Metadataset (USA - Multi-institutional): Machine learning ready nodule-level dataset with comprehensive feature engineering
- v1.0.0 (January 2026): Initial release with 13 curated datasets
- NSCLC-Radiomics (Netherlands)
- UniToChest (Italy)
- IMDCT (China - Multi-institutional)
- LNDb v4 (Portugal)
- BIMCV-R (Spain)
- LUNGx (USA)
- LIDC-IDRI (USA - Multi-institutional)
- LUNA25 (Netherlands - Multi-institutional)
- MIDRC-RICORD (USA - Multi-institutional)
- U-10 United-10 (Multi-national - 10 datasets)
- NLST-3D (USA - Multi-institutional)
- DLCS 2024 (USA - Duke University)
- DeepLesion-1kTest3D (USA - NIH Clinical Center)
- More updates coming soon...
⭐ If you find this resource helpful, please consider starring this repository!